



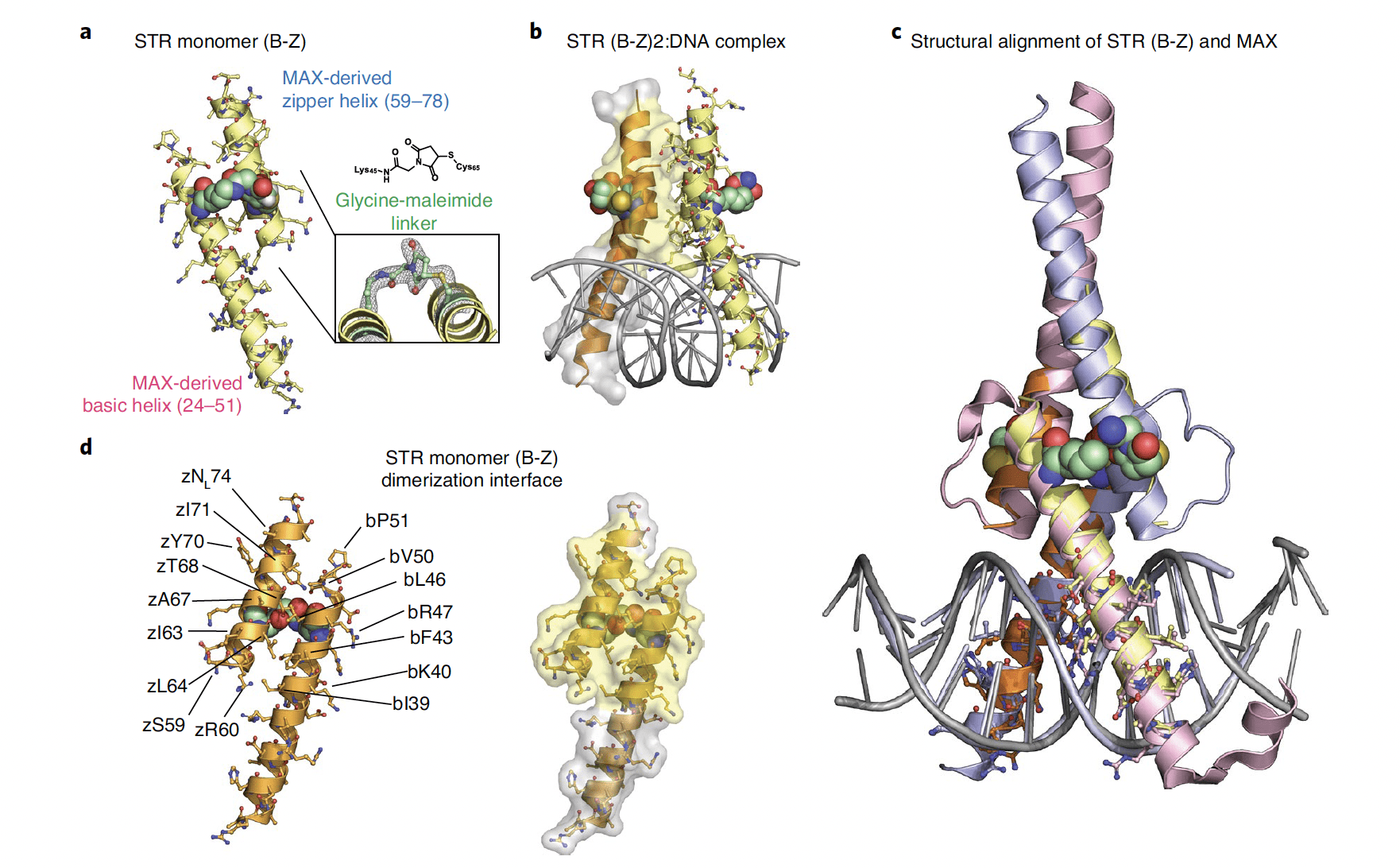

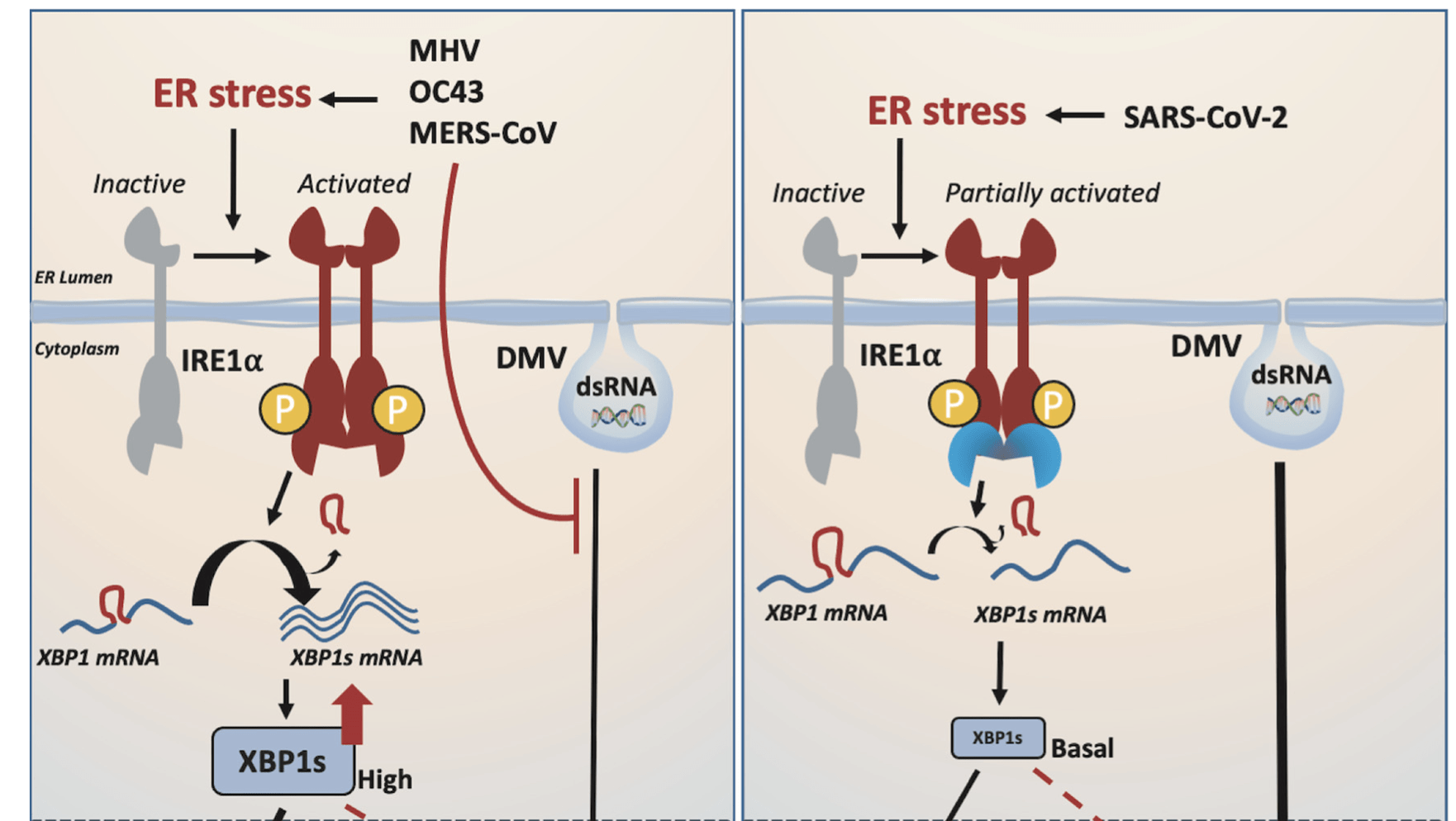
SARS-CoV-2 diverges from other betacoronaviruses in only partially activating the IRE1a/XBP1 ER stress pathway in human lung-derived cells.
Nguyen LC, Renner DM, Silva D, Yang D, Medina KM, Nicolaescu V, Gula H, Drayman N, Valdespino A, Mohamed A, Dann C, Wannemo K, Robinson-Mailman L, Gonzalez A, Stock L, Cao M, Qiao Z, Moellering RE, Tay S, Randall G, Beers MF, Rosner MR, Oakes SA & Weiss SR. mBio; 2022, 13(5):e0241522.
Diels-Alder Cycloadditions for Peptide Macrocycle Formation
Montgomery JE & Moellering RE. Methods in Mol Biol.; 2022, 2371: 159-74.

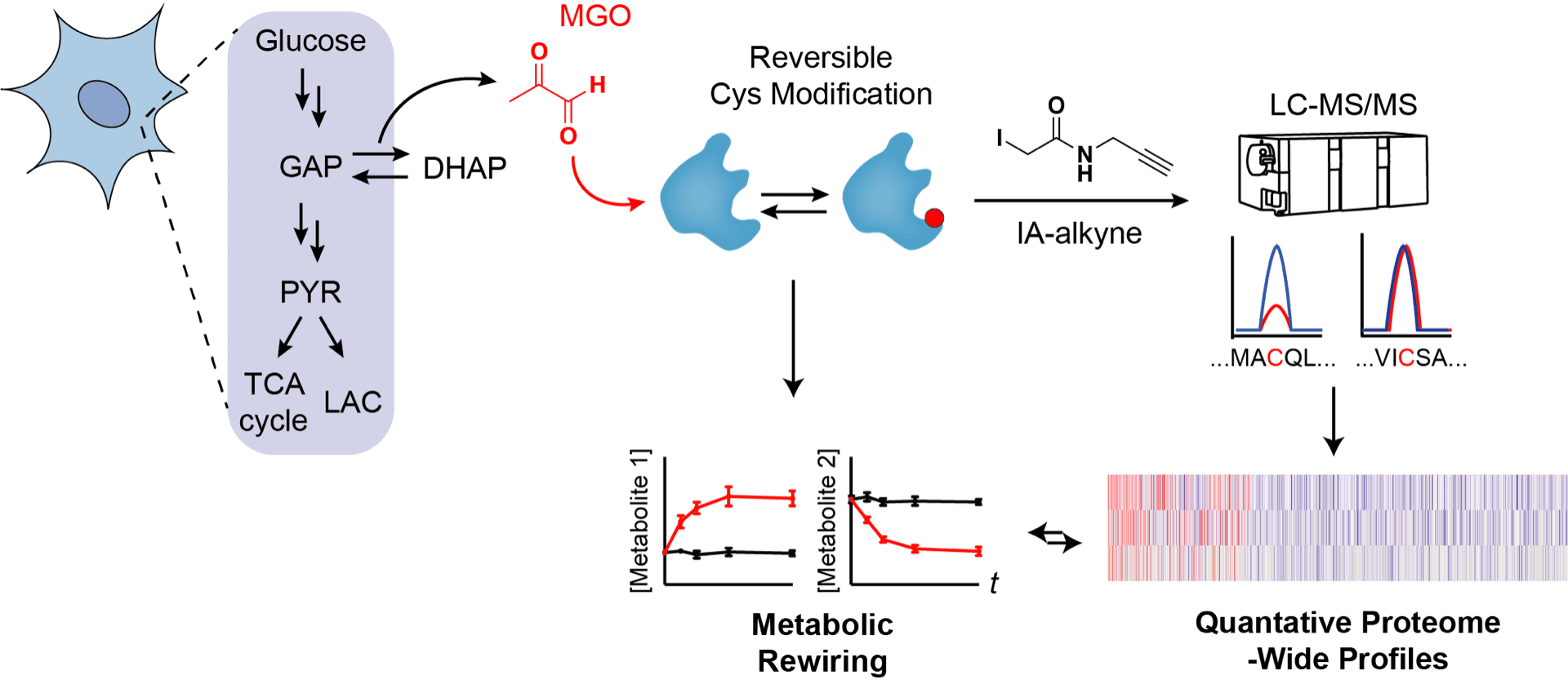
Methylglyoxal Forms Diverse Mercaptomethylimidazole Crosslinks with Thiol and Guanidine Pairs in Endogenous Metabolites and Proteins
Coukos JS & Moellering RE. ACS Chem. Biol.; 2021, 16(11): 2453-61.

Protein Targeting with SAF(er) Electrophiles
Speltz TE & Moellering RE. Nat. Chem.; 2020, 12(10): 885-7. (invited News & Views)
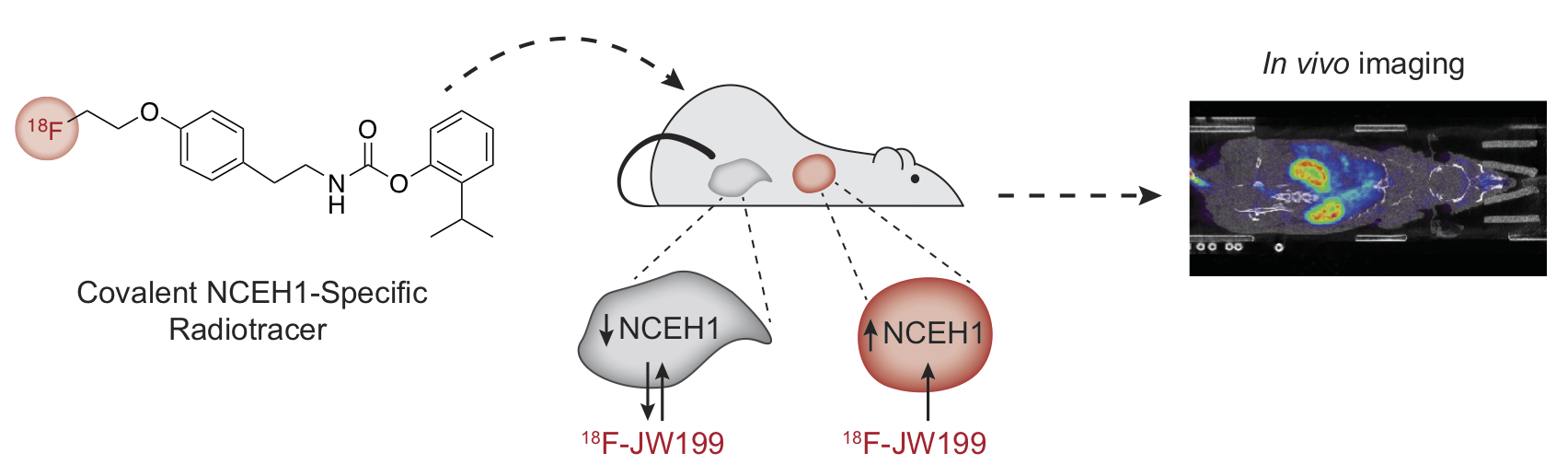
In vivo imaging of the tumor-associated enzyme NCEH1 with a covalent PET probe.
Chang JW, Bhuiyan M, Tsai HM, Zhang HJ, Li G, Fathi S, McCutcheon DC, Leoni L, Freifelder R, Chen CT & Moellering RE. Angew. Chem. Int. Ed.; 2020, 59(35): 15161-5.
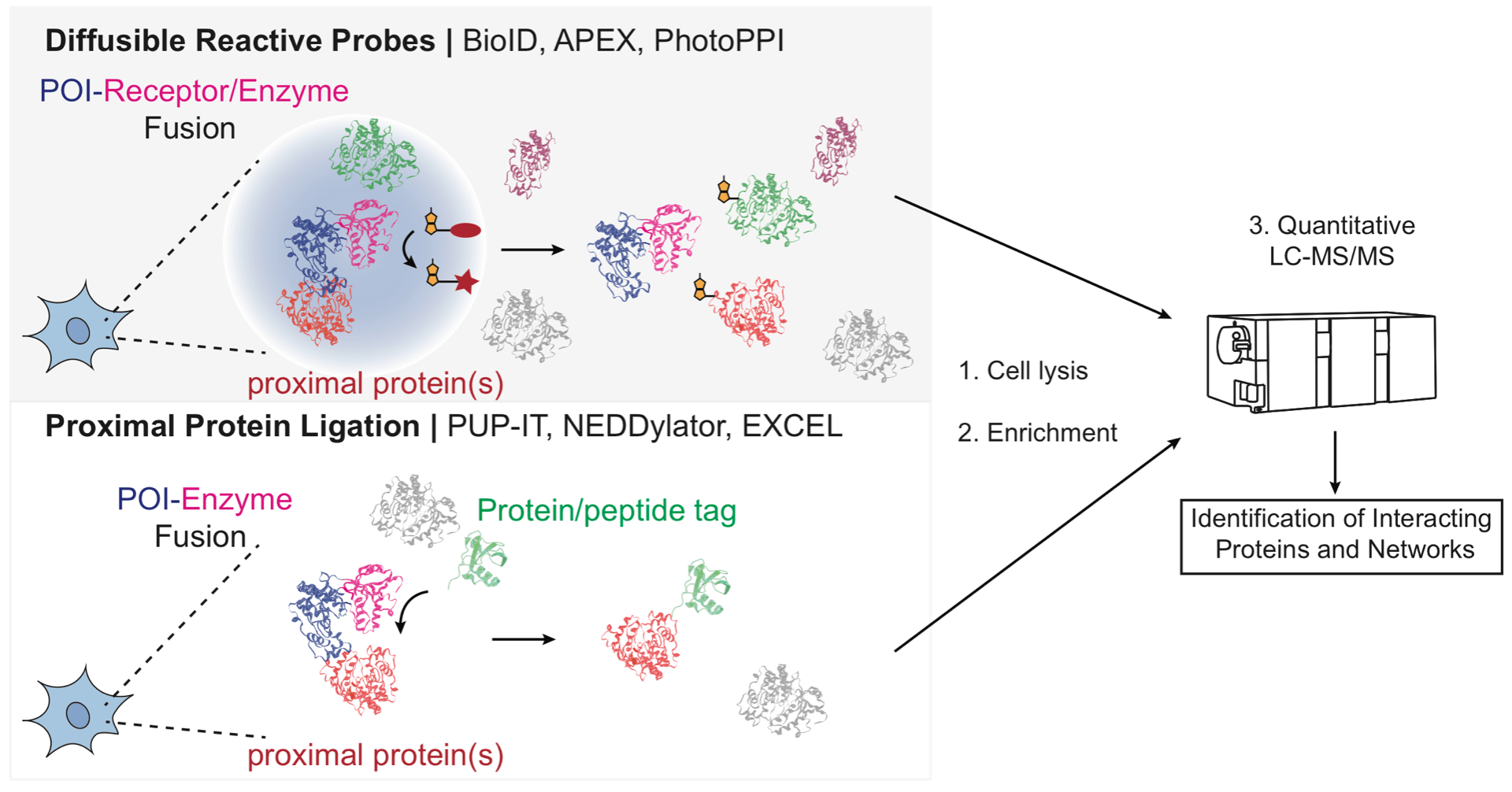
Interaction profiling methods to map protein and pathway targets of bioactive ligands.
Huang JX, Coukos JS & Moellering RE. Curr. Opin. Chem. Biol.; 2020, 54: 76-84.

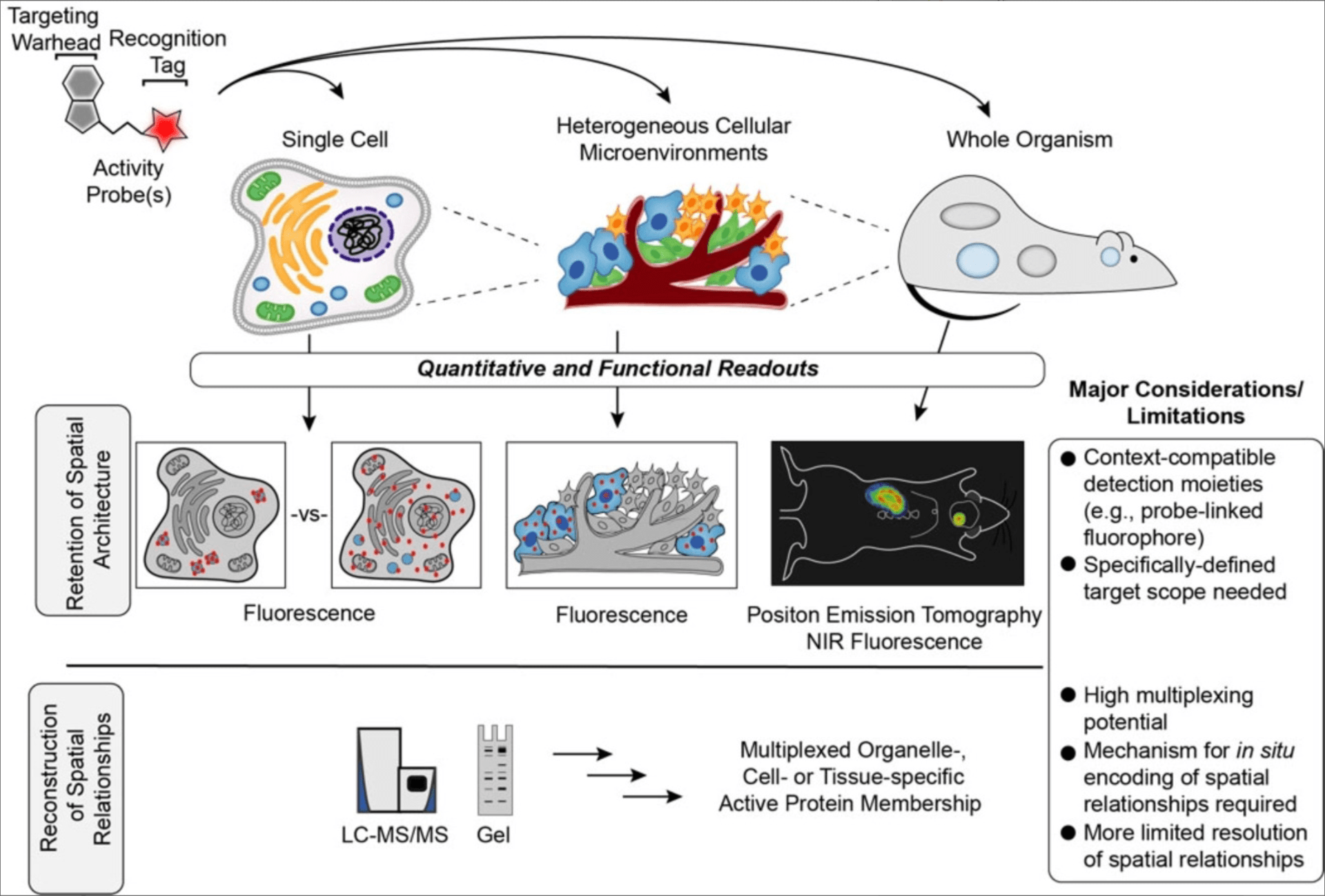
Tracking the ties that bind: emerging technologies to capture biomolecular interactions in native environments.
Moellering RE. Curr. Opin. Chem. Biol.; 2020, 54: A1-4.
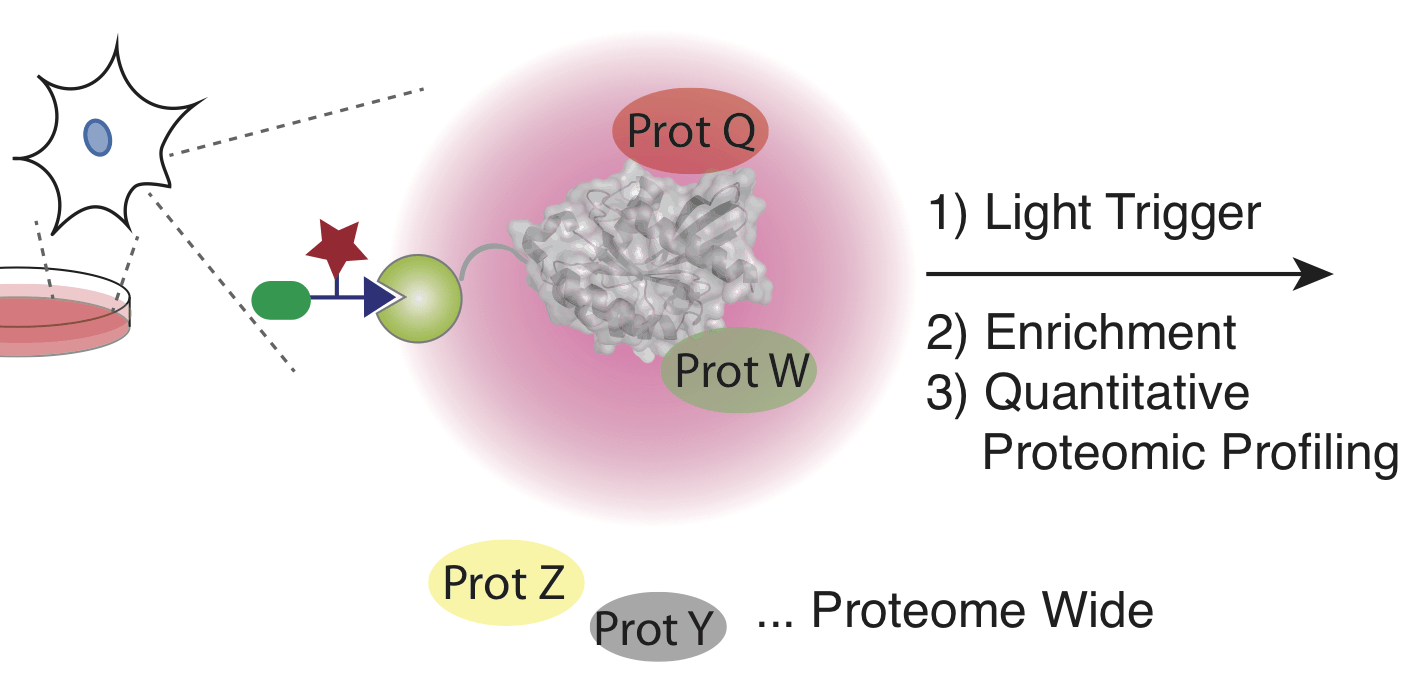
Photoproximity profiling of protein-protein interactions in cells.
McCutcheon DC, Lee G, Carlos AJ, Montgomery JE & Moellering RE. J. Am. Chem. Soc.; 2020, 142: 146-53.
bioRxiv preprint: https://www.biorxiv.org/content/10.1101/833384v1
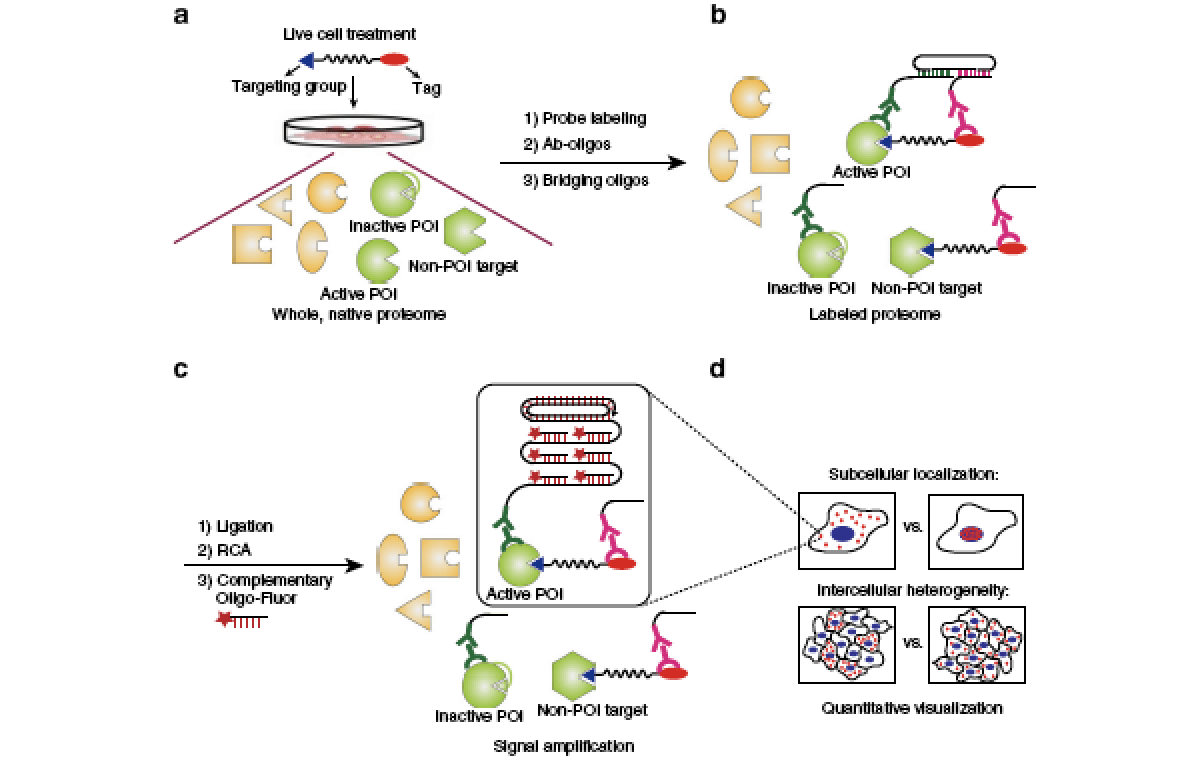
Ultrasensitive, multiplexed chemoproteomic profiling with soluble activity-dependent proximity ligation.
Li G, Eckert MA, Chang JW, Montgomery JE, Chryplewicz A, Lengyel E & Moellering RE. Proc. Natl. Acad. Sci. U.S.A.; 2019, 116(43): 21493-500.
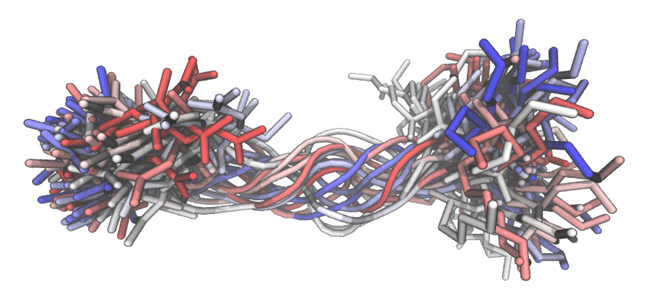
Versatile peptide macrocyclization with Diels-Alder cycloadditions.
Montgomery JE, Donnelly J, Fanning S, Speltz T, Shangguan X, Coukos J, Greene G & Moellering RE. J. Am. Chem. Soc., 2019; 141(41): 16374-81.
Faculty of 1000

High throughput discovery of functional protein modifications by hotspot thermal profiling.
Huang JX, Lee G, Cavanaugh KE, Chang JW, Gardel ML & Moellering RE. Nat. Methods, 2019, 16(9):894-901.
Open Access Protocol via ProtocolExchange: Here.

No bones about it: Small molecules for bone regeneration.
Chang JW & Moellering RE. Cell Chem. Biol., 2019, 26: 911-2. (editorial Preview).
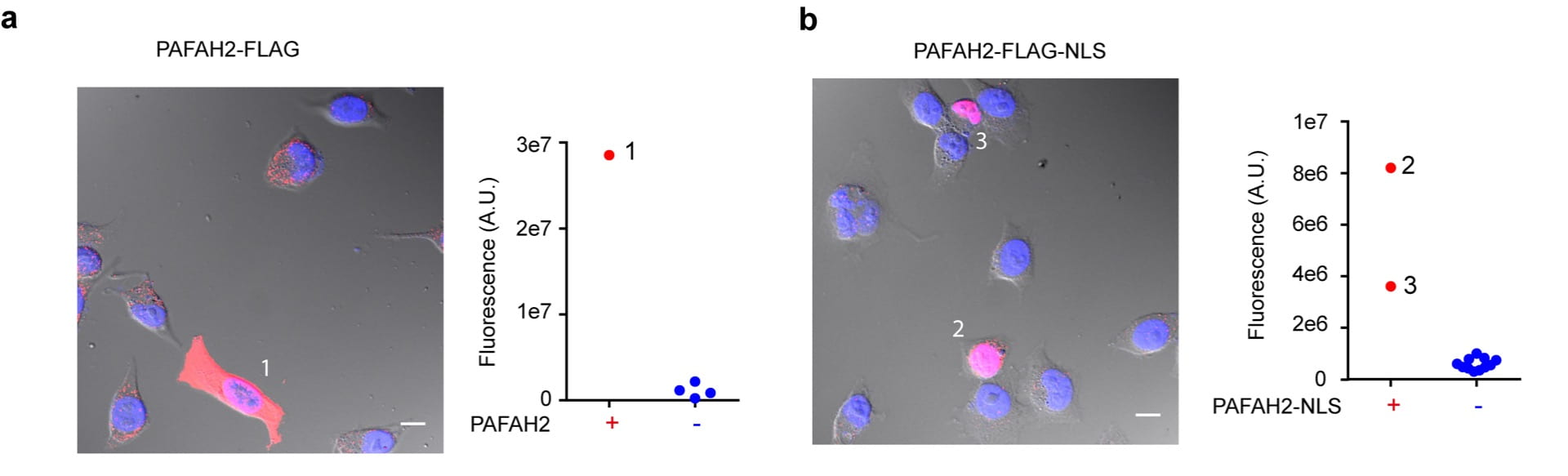
Chemical probes for spatially-resolved measurement of active enzymes in single cells.
Li G & Moellering RE. Methods Enzymol., 2019, 628: 243-262.
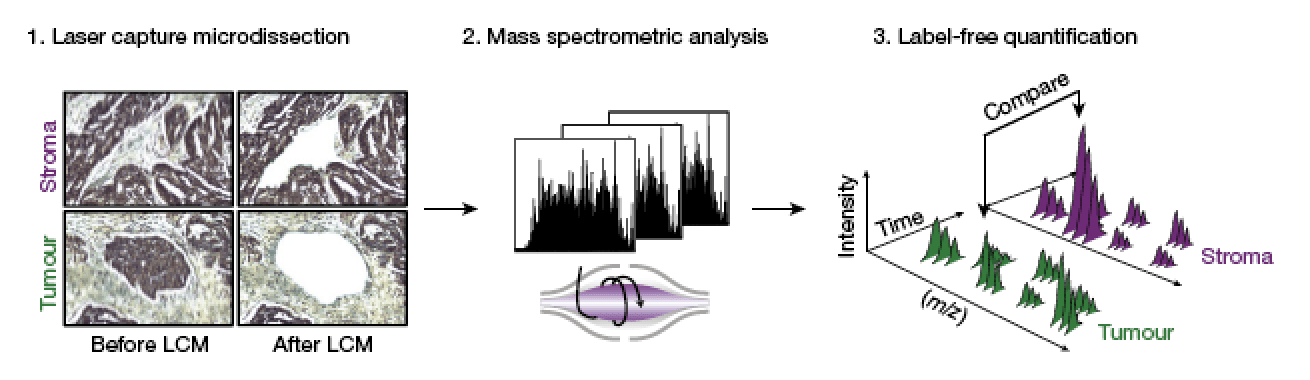
Proteomics reveals NNMT as a master metabolic regulator of cancer associated fibroblasts.
Eckert MA, Coscia F, Chryplewicz A, Chang JW, Hernandez KM, Pan S, Tienda S, Nahotko DA, Li G, Blazenovic I, Lastra RR, Curtis M, Yamada SD, Perets R, McGregor SM, Andrade J, Fiehn O, Moellering RE, Mann M & Lengyel E. Nature, 2019, 569(7758): 723-8.
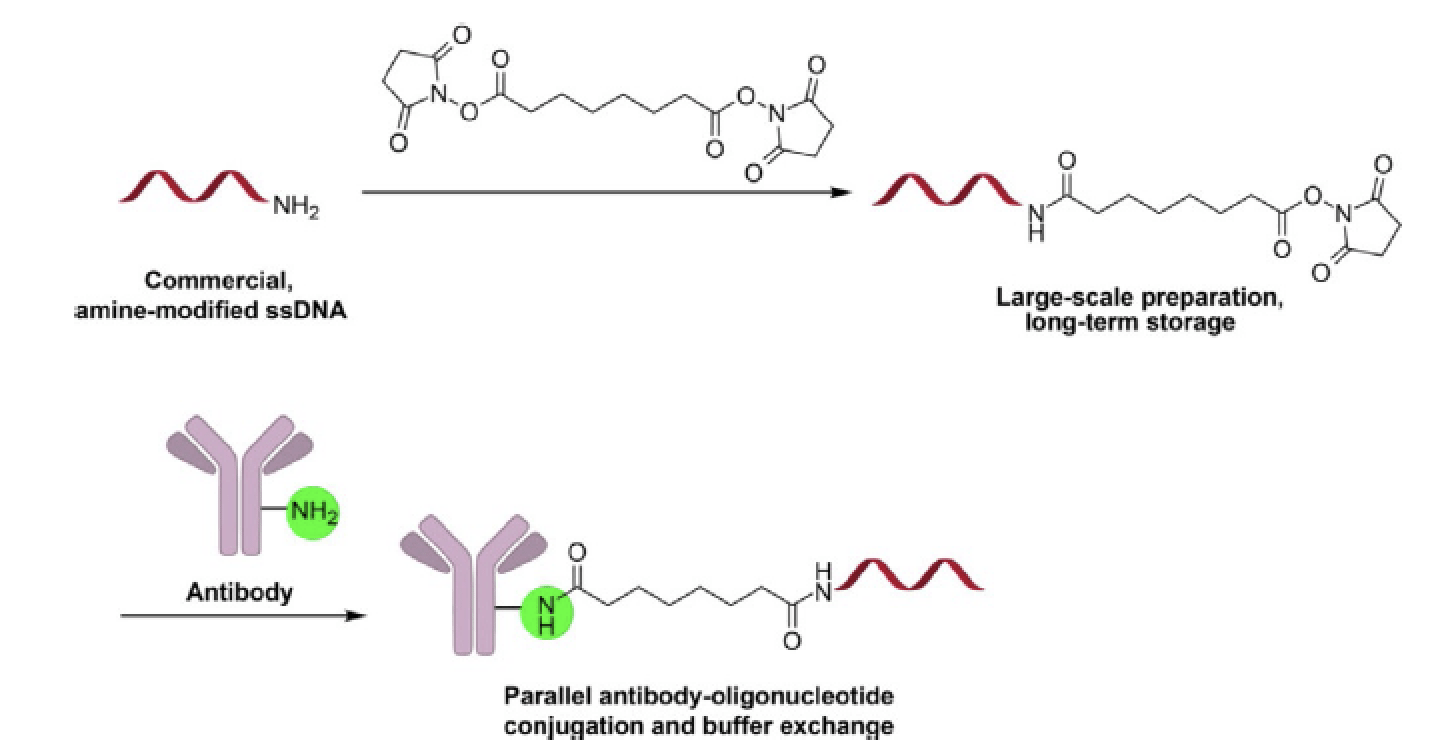
A concise, modular antibody-oligonucleotide conjugation strategy based on disuccinimidyl ester activation chemistry.
Li G & Moellering RE. ChemBioChem, 2019, 20(12): 1599-1605.
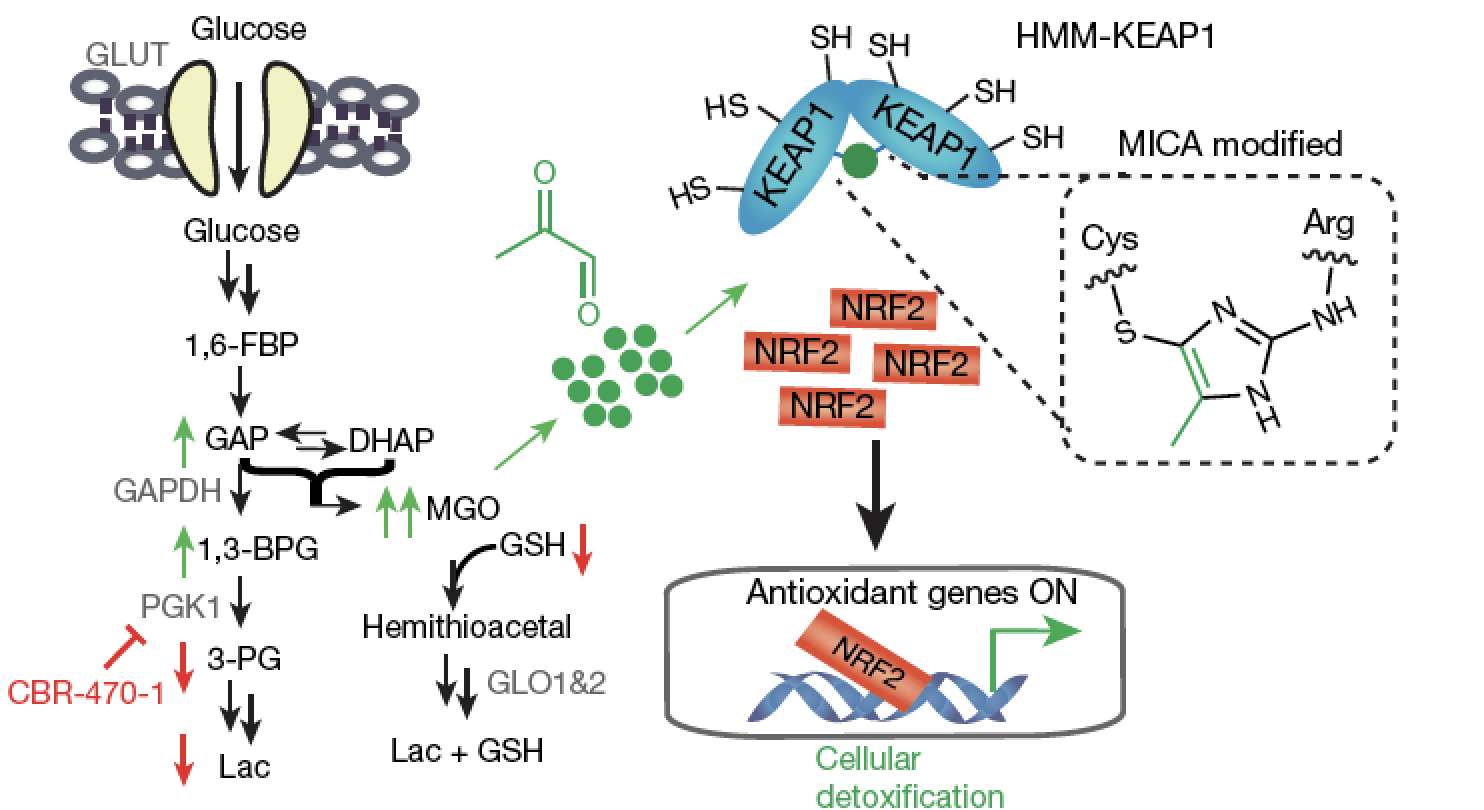
A metabolite-derived protein modification integrates glycolysis with KEAP1-NRF2 signaling.
Bollong MJ#, Lee G#, Coukos JS, Yun H, Zambaldo C, Chang JW, Chin EN, Ahmad I, Chatterjee AK, Lairson LL, Schultz PG & Moellering RE. Nature, 2018; 562(7728): 600-4.
Research Highlight: Nature Chemical Biology.
Faculty of 1000.
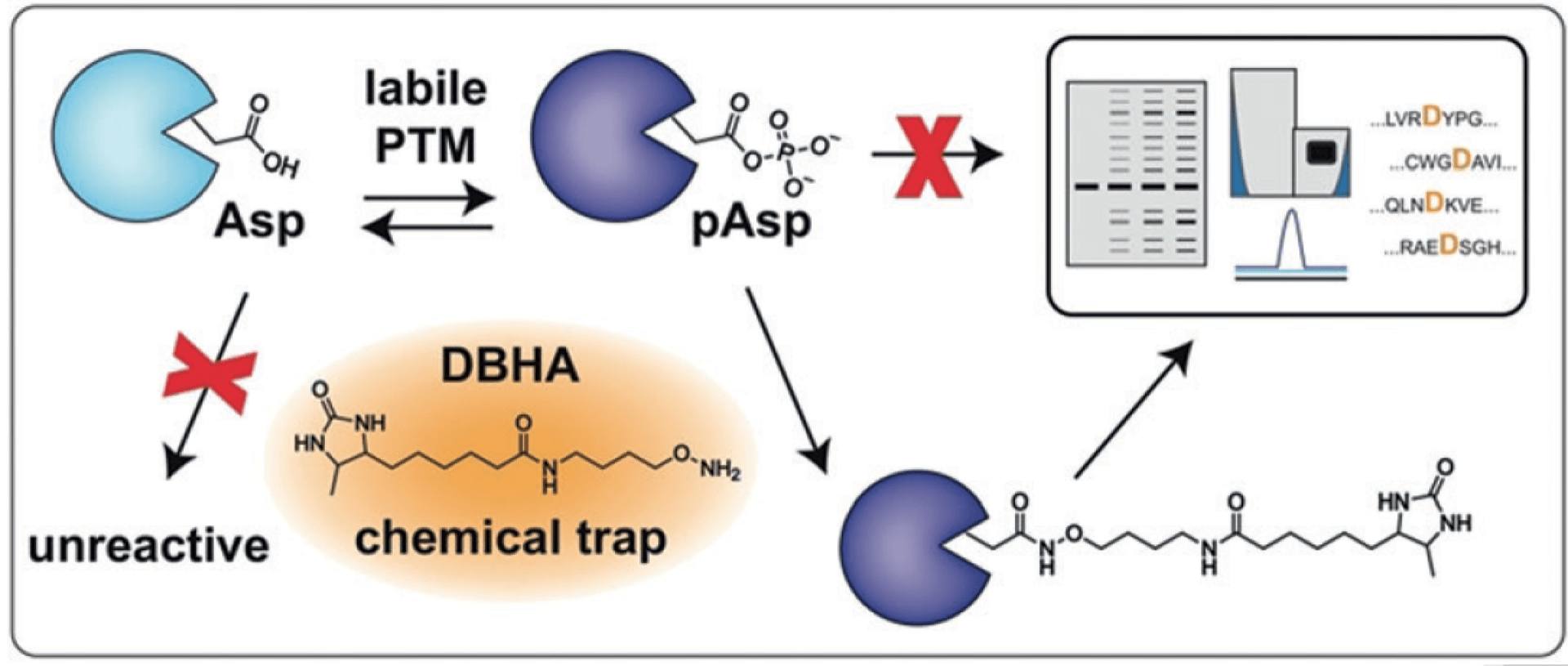
Chemoproteomic profiling of phosphoaspartate modifications in prokaryotes.
Chang JW, Lee G, Montgomery JE & Moellering RE. Angew. Chem. Int. Ed., 2018; 57(48): 15712-6.
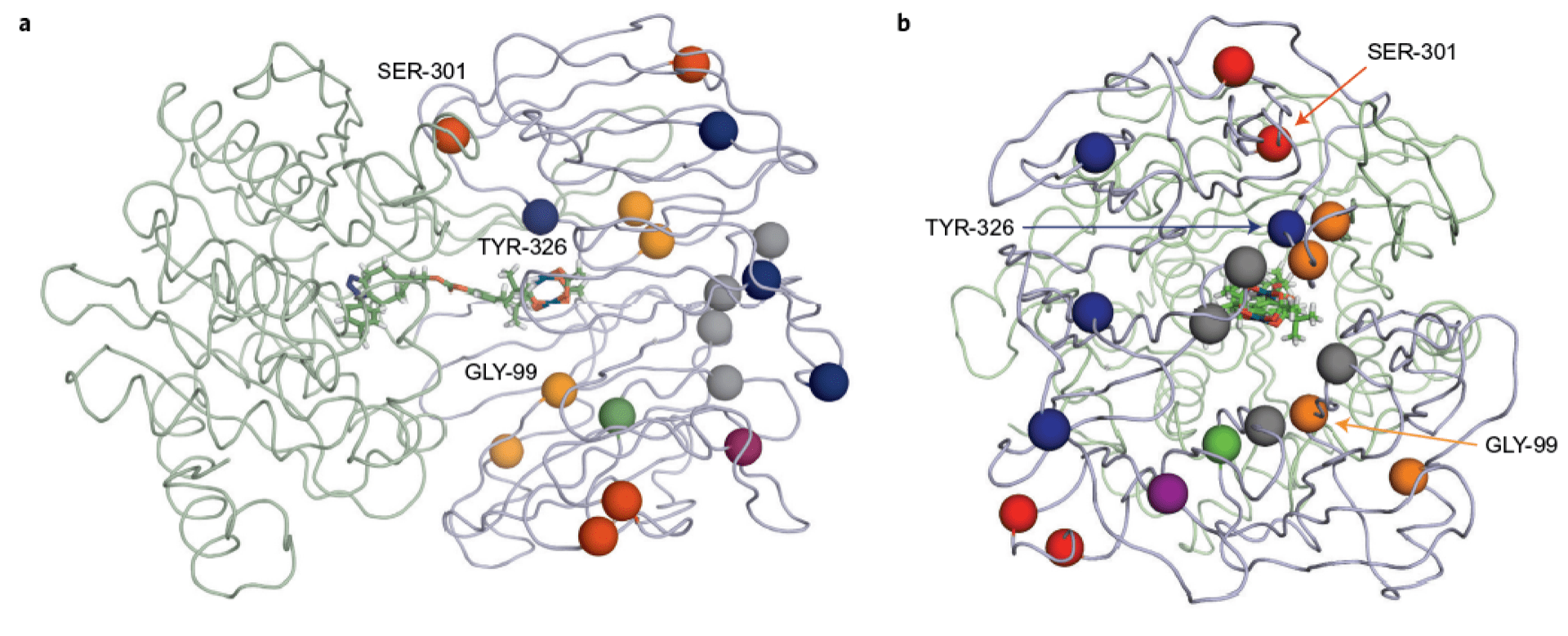
Evolving artificial metalloenzymes via random mutagenesis.
Yang H, Swartz AM, Ellis-Guardiola K, Park HJ, Upp DM, Lee G, Zhang C, Srivastava P, Moellering RE & Lewis JC. Nat. Chem., 2018; 10(3): 318-24.
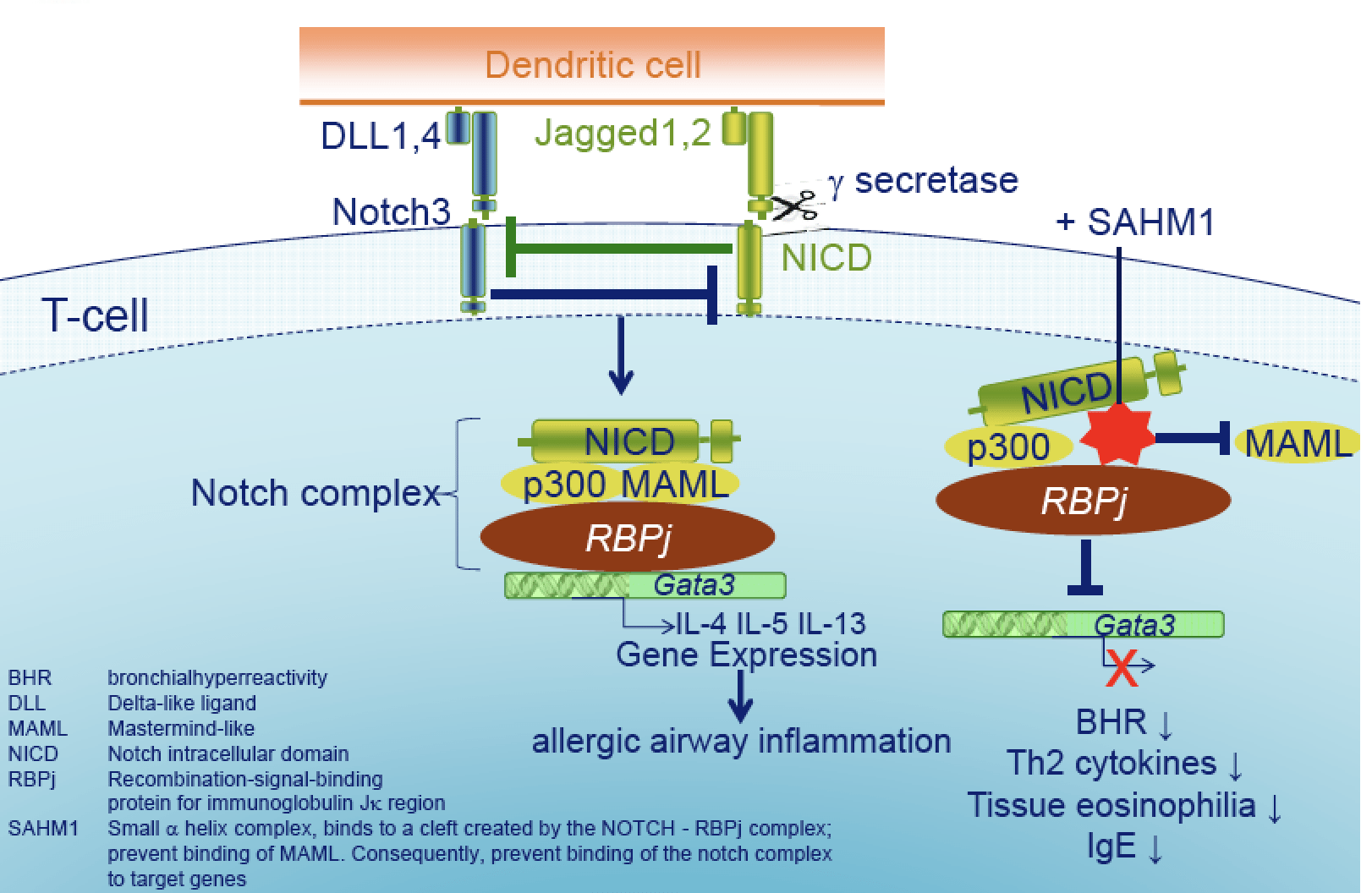
The Notch pathway inhibitor stapled alpha-helical peptide derived from mastermind-like 1 (SAHM1) abrogates the hallmarks of asthma.
KleinJan A, Nimwegan MV, Tindemans I, Bruijn M, Bergen I, Montgomery JE, Moellering RE, Hoogsteden H, Boon L, Amsen A & Hendriks RW. J. Allergy Clin. Immunol., 2018; 142(1): 76-85.

An activity-dependent proximity ligation platform for spatially resolved quantification of active enzymes in single cells.
Li G, Montgomery JE, Eckert MA, Chang JW, Tienda SM, Lengyel E & Moellering RE. Nat. Commun., 2017; 8(1): 1775.
Methods in Brief: Nature Methods.
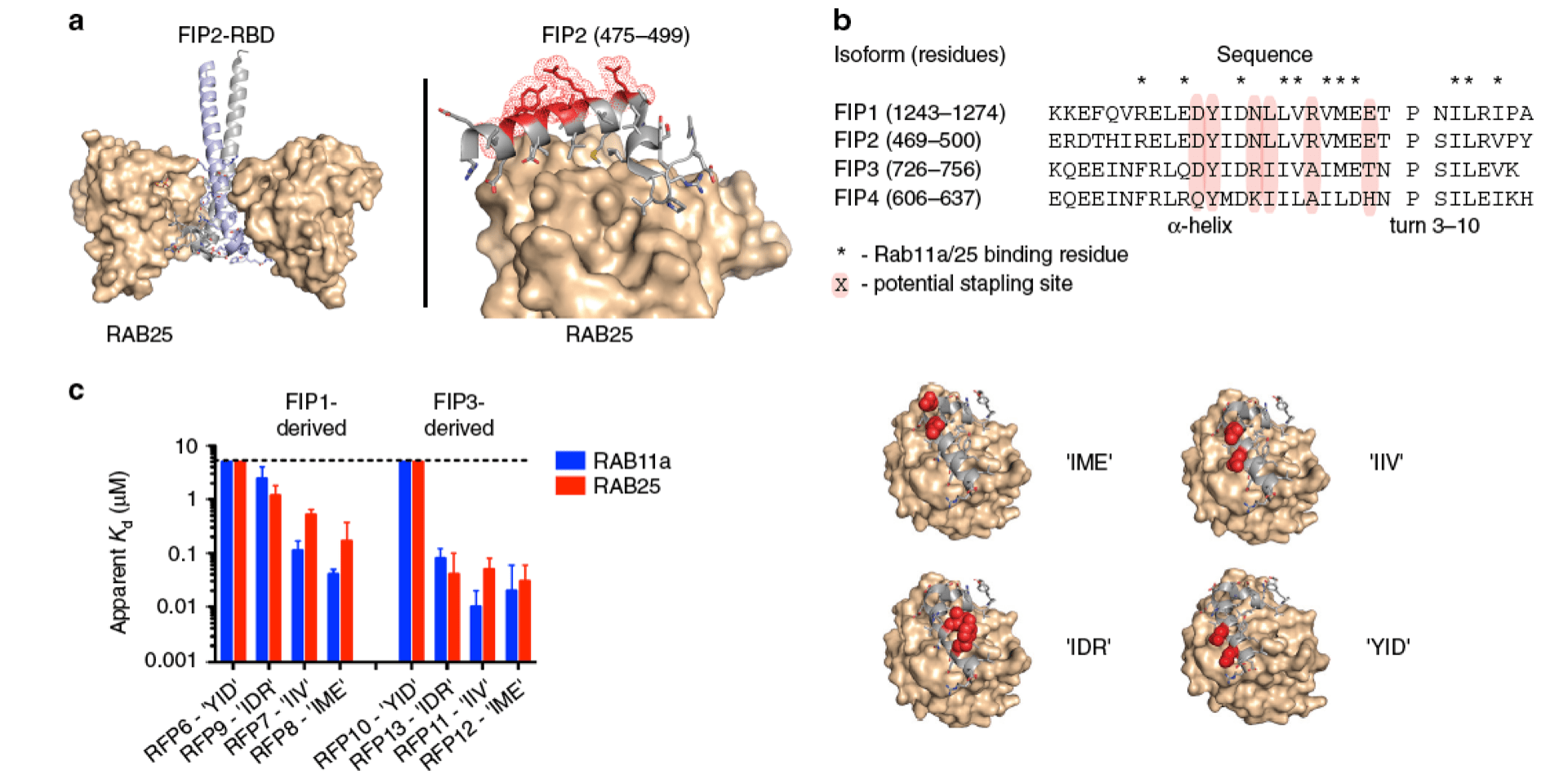
Stapled peptide inhibitors of RAB25 target context-specifc phenotypes in cancer.
Mitra S, Montgomery JE, Kolar MJ, Li G, Jeong KJ, Peng B, Verdine GL, Mills GB & Moellering RE. Nat. Commun., 2017; 8(1): 660.

Profiling reactive metabolites via chemical trapping and targeted mass spectrometry.
Chang JW, Lee G, Coukos JS & Moellering RE. Anal. Chem., 2016; 88 (13): 6658-61.
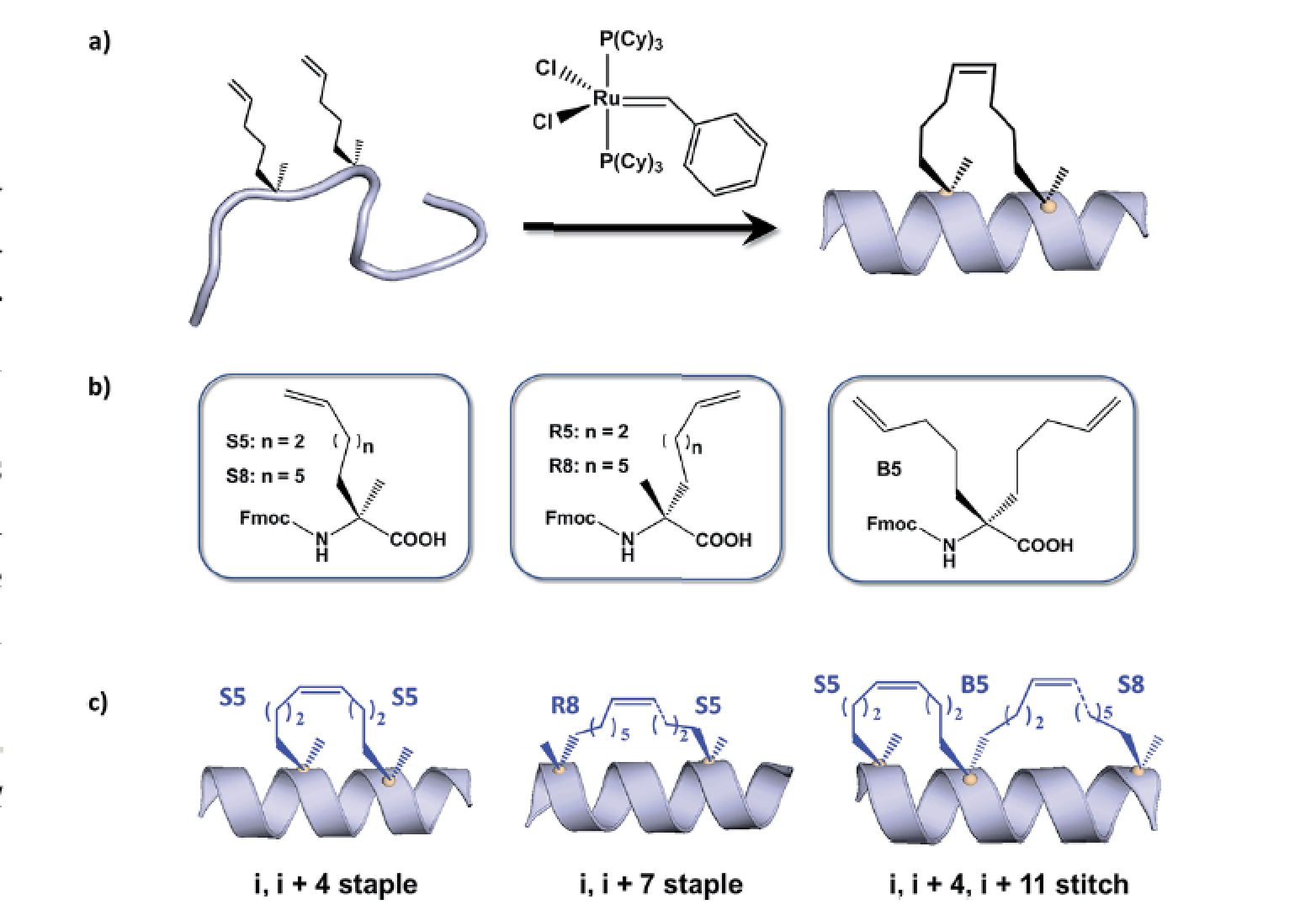
Towards understanding cell penetration by stapled peptides.
Chu Q#, Moellering RE#, Hilinski GJ, Kim YW, Grossmann TN, Yeh JT & Verdine GL. Med. Chem. Commun., 2015; 6, 111-9.
# – co-first author
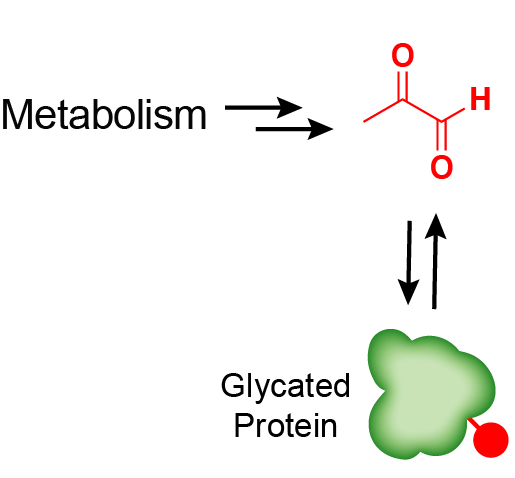
Functional lysine modification by an intrinsically reactive primary glycolytic metabolite.
Moellering RE* & Cravatt BF*. Science, 2013; 341 (6145): 549-53. *-co-corresponding author.
Editor’s Choice in Science Signaling.
Research Highlight in Nature Chemical Biology.
Highlighted in Faculty of 1000.
Spotlight in ACS Chemical Biology.
Inhibition of oncogenic WNT signaling through direct targeting of ß-catenin.
Grossmann TN, Yeh JTH, Bowman BR, Chu Q, Moellering RE & Verdine GL. Proc. Natl. Acad. Sci., 2012; 109(44): 17942-7.
How chemoproteomics can enable drug discovery and development.
Moellering RE & Cravatt BF. Chem. Biol., 2012; 19(1): 11-22.
An activity-based imaging probe for the integral membrane serine hydrolase KIAA1363.
Chang JW#, Moellering RE# & Cravatt BF. Angew. Chem. Int. Ed., 2012; 51: 966-70. # – co-first author
Direct inhibition of the NOTCH transcription factor complex.
Moellering RE, Cornejo M, Davis TN, Del Bianco C, Aster JC, Blacklow SC, Kung AL, Gilliland DG, Verdine GL & Bradner JE. Nature, 2009; 462: 182-8.
News and Views Nature.
News of the Week in C&EN News.
Highlighted in Science Signaling.
Preview in Developmental Cell.
Spotlight in ACS Chemical Biology.
Highlighted in Faculty of 1000.
News Highlight BBC.
Acid treatment of melanoma cells selects for invasive phenotypes.
Moellering RE, Black K, Krishnamurthy C, Baggett BK, Stafford P, Rain M, Gatenby RA & Gillies RJ. Clin. Exp. Metastasis, 2008; 25: 411-25.

